Sulfatase family S1
Family S1 comprises the vast majority of the
sulfatases. They catalyze the removal of sulfate ester groups
according a hydrolytic mechanism (EC 3.1.6.-sulfuric ester
hydrolases; EC 3.10.1.- sulfamidases).
Family S1 sulfatases containing a unique catalytic residue, the
Cα-formylglycine (FGly), which is posttranslationally generated from
a conserved cysteine or serine. The posttranslational modification
occurs when the polypeptide chain is still unfolded and is directed
by a conserved N-terminal [C/S]XPXR motif (Schmidt et al. 1995; Miech
et al. 1998). These S1 sulfatases adopt a similar fold comprising two
(alpha/beta) domains, a large N-terminal domain containing the
catalytic pocket and a smaller C-terminal domain. The active site
encompasses the catalytic nucleophile formylglycine (Cys69, human
ARSA numbering, family S1-1) and nine additional conserved residues
(Lukatela et al. 1998): (i) four acidic/polar residues (Asp29, Asp30,
Asp281, and Asn282) coordinating a calcium ion which binds and
activates the sulfate group of the substrate (ii) five basic amino
acids (Arg73, Lys123, His125, His229, and Lys302) stabilizing the
formylglycine and/or binding the sulfate group.
Subfamilies (111)
EC activities found in subfamilies
| Subfamily |
EC Number(s) |
Description |
PubMed IDs |
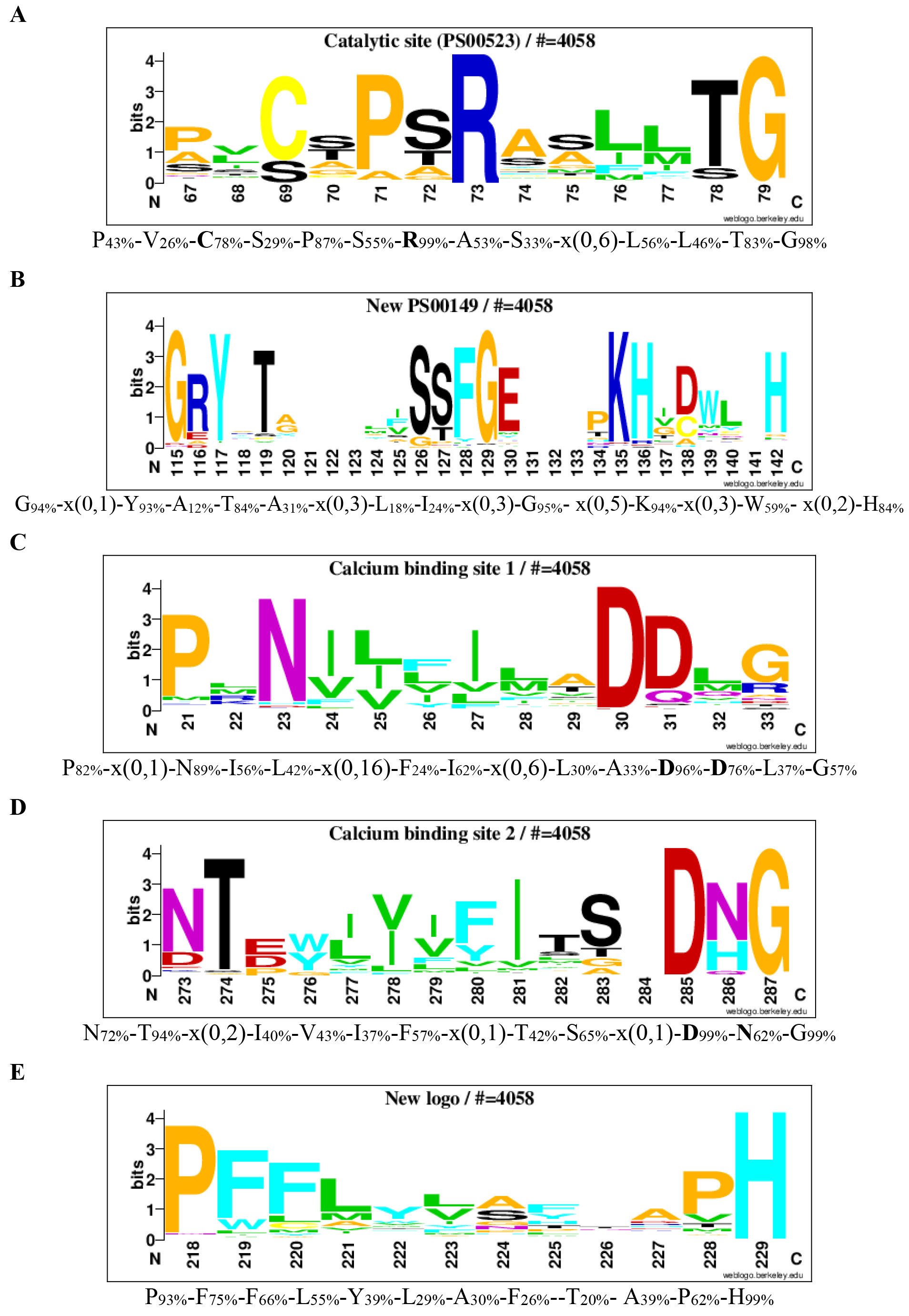
The consensus sequence logoplots of family S1 (Barbeyron et al. 2016)
Logos of conserved consensus sequences identified in the global alignment of formylglycine-dependent sulfatases (S1 family).
Logos of conserved consensus sequences were identified from 4058 aligned S1 sulfatases. The logo sequence of the catalytic site that corresponds to the PROSITE signature PS00523, is shown in A. The logo sequence of PROSITE signature PS00149 is shown in B. The two logo sequences of calcium binding are shown in C and D. A logo sequence from a conserved supplementary consensus sequences is shown in E. The numbers below the logo sequences indicate, at the first position, the corresponding position in reference sequence (AtsA, P51691). The corresponding consensus sequences in multi-alignment are shown below the logo sequences. The percentages in subscript are the percentages of sequences, where the amino acid is conserved in alignment. Catalytic amino acids and residues involved in calcium ion binding are in bold.
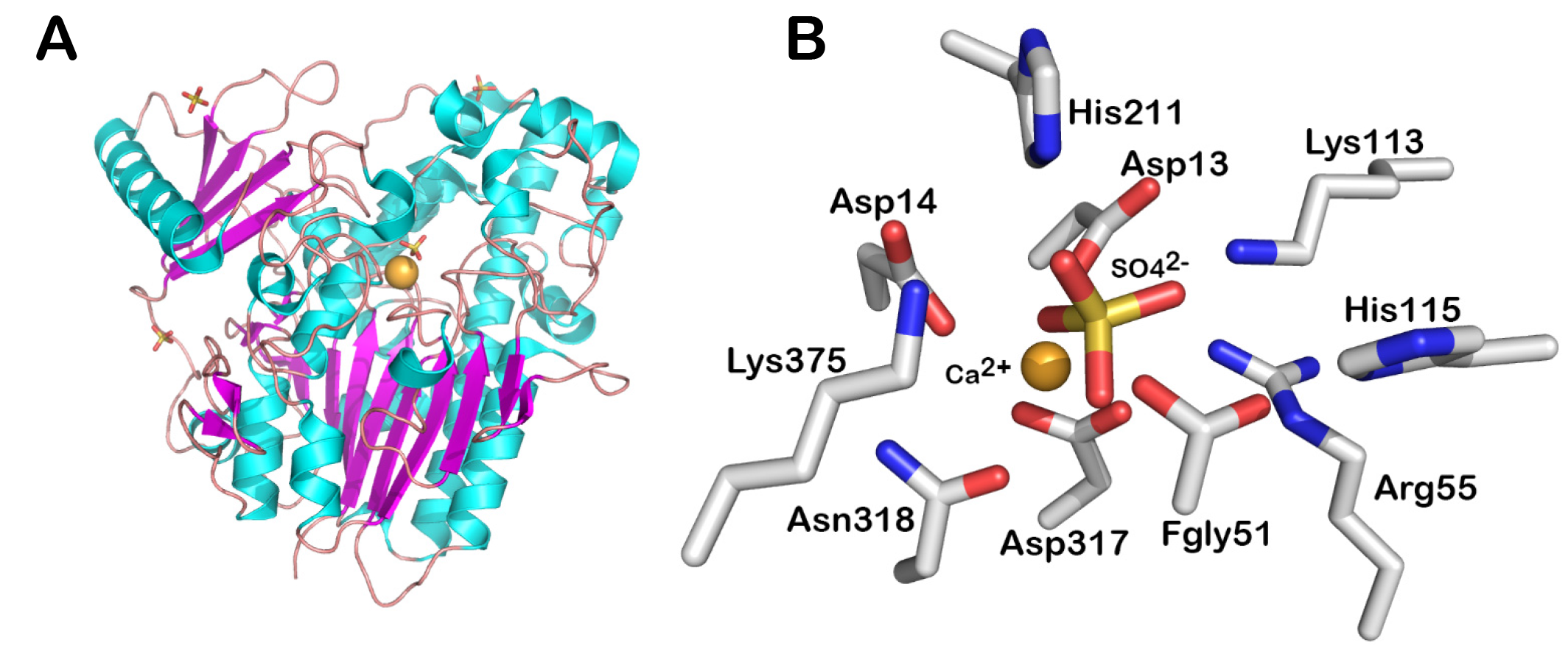
Fold and active site of a S1 family representative
Fold (A) and active site (B) of the arylsulfatase AtsA from Pseudomonas aeruginosa PAO1 (PDB code: 1HDH). The fold is shown in cartoon representation. The amino acids and ligands of the active site is shown in sticks. The cations are shown as spheres. The figures were made using PyMoL (Version 1.8 Schrödinger, LLC).
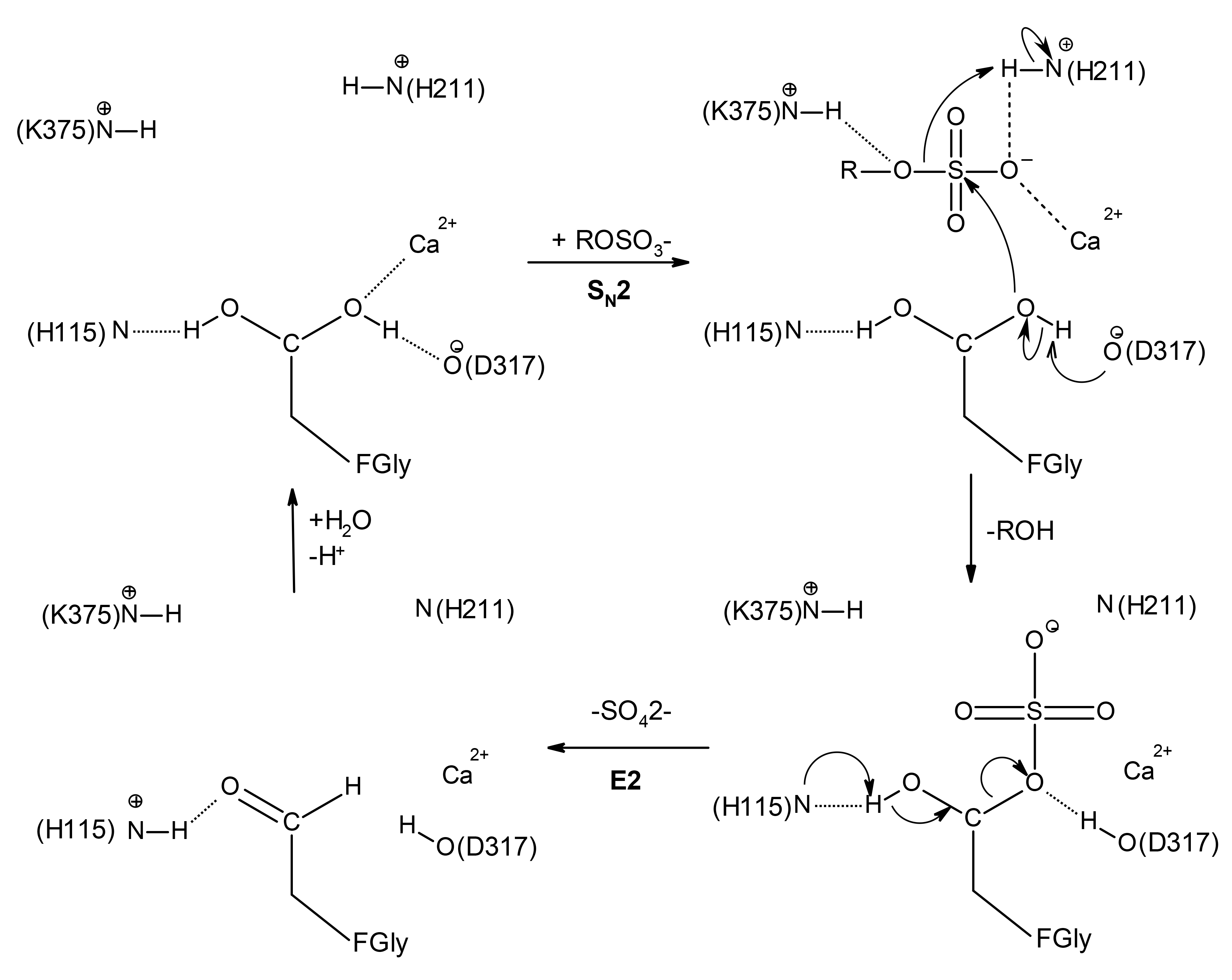
Favored catalytic mechanism of the S1 family sulfatases.
The numbering corresponds to the arylsulfatase AtsA from Pseudomonas aeruginosa PAO1. Upon substrate binding, the formyglycine is activated for nucleophilic attack on sulfur by Asp317. The sulfoenzyme intermediate is formed, and desulfation most likely occurs by elimination from the remaining fGly-diol hydroxyl (E2), catalyzed by His115. This figure was adapted from the following references (Boltes et al, 2001, Structure; Appel and Bertozzi, 2015, ACS Chem Biol) and prepared with Accelrys Draw 4.2.
 Family details
Family details




